Any decompositional clustering algorithm can be made hierarchical by recursive application.
K-means-cluster (in S : set of vectors : k : integer)
{ let C[1] ... C[k] be a random partition of S into k parts;
repeat {
for i := 1 to k {
X[i] := centroid of C[i];
C[i] := empty
}
for j := 1 to N {
X[q] := the closest to S[j] of X[1] ... X[k]
add S[j] to C[q]
} }
until the change to C (or the change to X) is small enough.
}
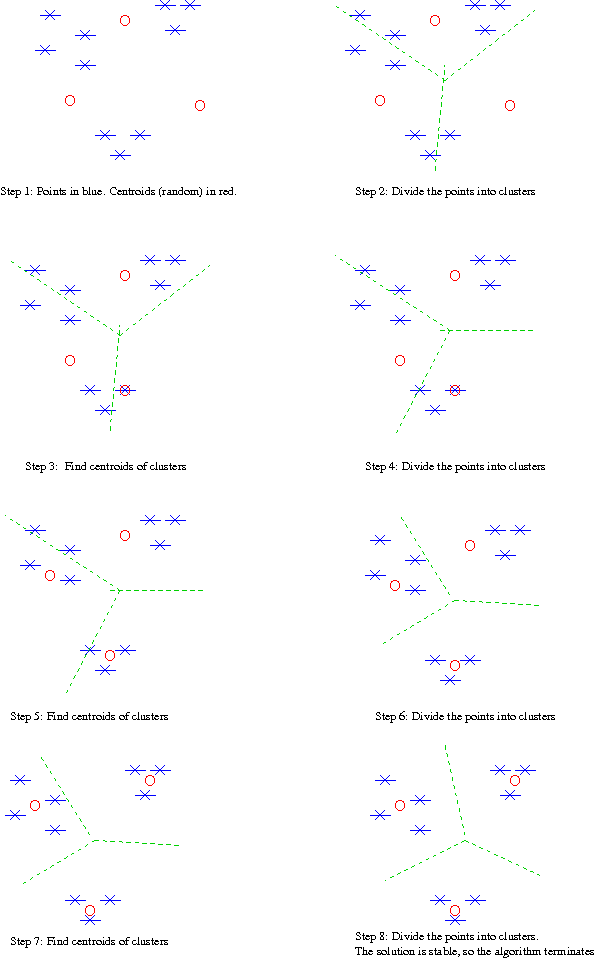
Example: (Note: I have started here with random points rather than a random partition.)

Features:
Sum of the squared distance steadily decreases over the iterations.
Hill-climbing algorithm.
Problems:

Variant 1: Update centroids incrementally. Claims to give better results.
Variant 2: Bisecting K-means algorithm:
for I := 1 to K-1 do {
pick a leaf cluster C to split;
for J := 1 to ITER do split C into two subclusters C1 and C2;
choose the best of the above splits and make it permanent
}
Generates a binary clustering hierarchy. Works better (so claimed) than
K-means.
Variant 3: Allow overlapping clusters: If distances from P to C1 and to C2 are close enough then put P in both clusters.
Full mean vector would get very long (= number of different words in all the documents.) Solution: truncate after first M terms. (Typically M=50 or 100.)
{ put every point in a cluster by itself
for I := 1 to N-1 do {
let C1,C2 be the most mergeable pair of clusters;
create C parent of C1, C2
} }
Various measures of "mergeable" used.
Characteristics
pick a starting point D in S;
CC = { { D } } } /* Set of clusters: Initially 1 cluster containing D */
for Di in S do {
C := the cluster in CC "closest" to Di
if similarity(C,Di) > threshhold
then add Di to C;
else add { Di } to CC;
}
Features: